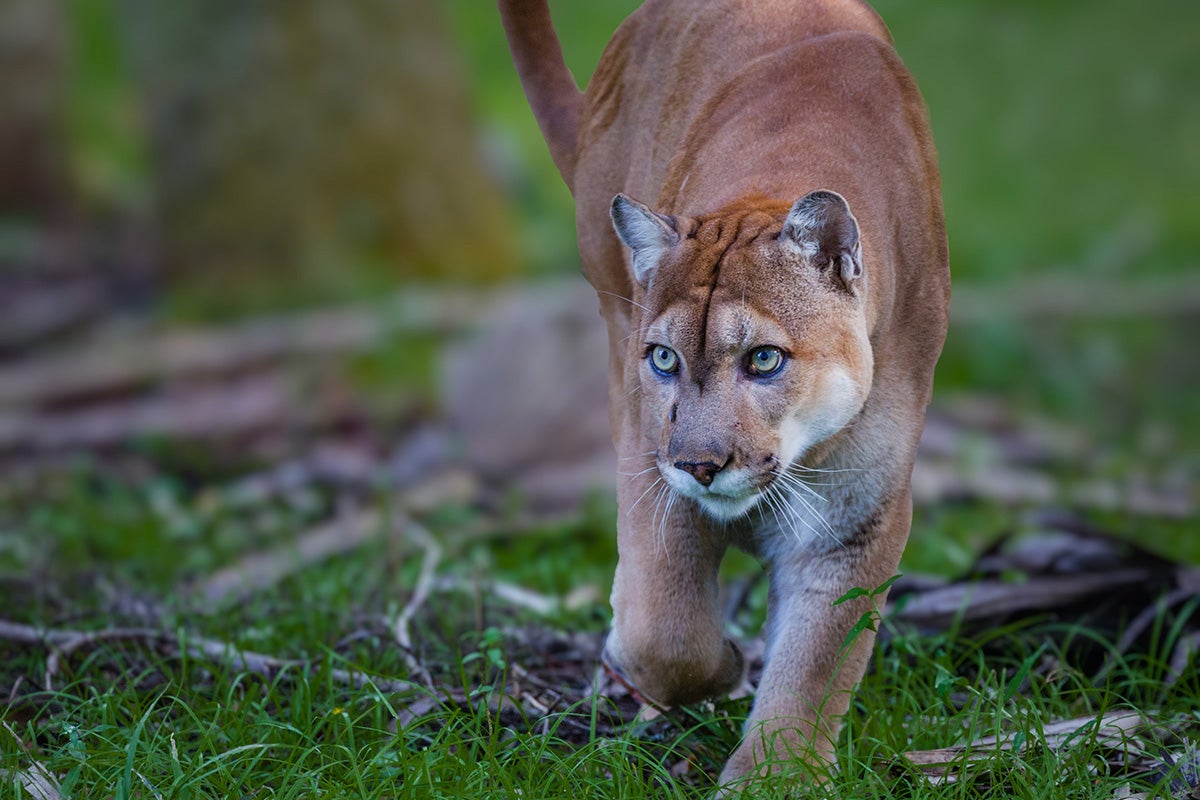
The introduction of Texas pumas to Florida in the 1990s as part of a genetic rescue may have helped save Florida panthers from extinction, but it also brought some harmful mutations with it along the way.
In a new study led by UCF, researchers show that nearly half of the harmful mutations found in recent Florida panthers came from the introduced Texas pumas, as well as from Central American pumas that were released into Everglades National Park in the 1950s and 1960s.
The findings, published recently in the Journal of Heredity, point to the importance of monitoring Florida panther population genetics as well as the need to genetically screen any future introductions.
The Florida panther’s population reached a dangerously low number of only 30 panthers in the 1970s and 1980s, which led to inbreeding and producing offspring with genetic problems, including heart defects, sexual defects such as cryptorchidism, and other diseases.
Genetic rescue efforts implemented in the 1990s brought pumas from Texas to Florida to breed with panthers in the Sunshine State. The process is known as genetic admixture.
The efforts appeared to help, with the Florida panther population rebounding to between 120 and 230 adults, according to estimates from the Florida Fish and Wildlife Conservation Commission, and reports of greater genetic variation now present in the population.
However, the new findings show this genetic influx had both good and bad mutations with it.
“There’s a genetic signal that harmful mutations from different source populations were introduced into the Florida panther population,” says the study’s lead author Alexander Ochoa, a postdoctoral scholar in UCF’s Department of Biology and Genomics and Bioinformatics research cluster.
“Although there’s still no evidence of these mutations emerging at the phenotypic level, we want to monitor the genetic health of the Florida panther because things could also go south pretty quickly, especially if their population remains small,” Ochoa says.
He says if researchers find an indication that the current fitness of the population is declining, then perhaps wildlife managers should consider future introductions and genetic rescue programs for Florida panthers.
“And if that’s the case, with the information we have in hand, we may be able to assess the genetic health of potential source populations, and pinpoint and select individuals that are best suited for these introductions,” Ochoa says. “Ideally, we would want to select individuals that carry smaller amounts of harmful, or deleterious, mutations for introductions.”
Surprising Discovery
An interesting discovery from the new research was the Central American roots of some of the deleterious genetic material.
Central American pumas were introduced to the Florida population, specifically to Everglades National Park, during poorly documented releases of captive pumas during the 1950s and 1960s.
As a result, Florida panthers from Everglades National Park in South Florida contain genetic ancestry from Central America, and like the introduction of pumas from Texas, brought good and bad mutations to the population.
The study found that 16% of the deleterious mutations present in Florida panthers are of exclusive Central American origin, whereas 33% of these mutations are of exclusive Texas origin. Another 4% of these deleterious mutations are of shared Central American and Texas origin.
For the study, researchers used whole-genome sequences to identify and compare levels of mutation load of panthers from Florida with no Texas or Central American ancestry (non-admixed); Everglades National Park panthers with Central American ancestry; Texas pumas; and panthers that had a mix of Florida and Texas ancestry.
They looked at proportions of harmful mutations and genotypes carrying these mutations.
The researchers found that, although genetic admixture increased Florida panther fitness by offsetting the expression of existing harmful mutations, admixed Florida panthers have now become carriers of many new harmful mutations.
“I was expecting that the genetic rescue program was in general beneficial to Florida panthers,” Ochoa says. “But at the same time one of the unexpected findings was that a reasonable amount of novel harmful mutations from different source populations were also introduced into the Florida panther population. And we need to monitor these mutations in current Florida panthers because that’s something that Florida panthers did not have before.”
Next Steps
Assistant Professor of Biology and study co-author Robert Fitak helped lead the research. He says the Florida panther has been a generally positive conservation success story, and that it now can also serve as a model to inform genetic rescues in other species.
“We know that in the future many more species are going to become endangered, suffer from severe inbreeding, and have to be managed by conservation actions such as genetic rescue,” Fitak says. “Our results will help us understand what the potential negative genetic consequences of these actions will be, or what we need to be prepared for.”
For Florida panthers, Fitak says next steps should include sequencing entire genomes of more individuals to explore the distribution of harmful genetic traits in the population.
“Until recently, most studies of wildlife genomes have focused on their adaptations, in other words, those traits that are beneficial and specific to the species,” Fitak says. “Now, we are flipping the question around and asking what about the bad, or ‘deleterious,’ traits?”
“We know that the genetic mixing has overall been a success for Florida panthers, but we need to be aware of any deleterious genetic stowaways that are tagging along for the ride,” he says.
Study co-authors were Dave Onorato with the Florida Fish and Wildlife Conservation Commission Fish and Wildlife Research Institute; Melody Roelke-Parker with Frederick National Laboratory for Cancer Research, Leidos Biomedical Research, Inc.; and Melanie Culver with the University of Arizona.
The work was supported by UCF’s Preeminent Postdoctoral Program.
Ochoa received his doctoral degree in natural resources from the University of Arizona and joined UCF in 2020.
Fitak received his doctorate in genetics from the University of Arizona and his bachelor’s in molecular genetics from The Ohio State University. Before joining UCF in 2019, he worked as a postdoctoral researcher at the Institute for Population Genetics in Vienna, Austria, and at Duke University. He is a member of UCF’s Genomics and Bioinformatics research cluster.
Study title: Give and Take: Effects of Genetic Admixture on Mutation Load in Endangered Florida Panthers