Most are familiar with the field of genomics in relation to ancestry mapping — such as 23andMe and Ancestry.com — uncovering relatives and identifying genetic risk factors for disease. However, genomics has also widely been applied to tracking infectious disease through sequencing the genomes of bacteria, parasites and viruses.
Now, it is helping us to better understand how to combat the coronavirus.
Assistant Professor Taj Azarian, an infectious-disease epidemiologist, is heading up genomic surveillance of SARS-CoV-2 at UCF ahead of the university’s plans for a full return to face-to-face instruction in the fall.
“One component to help us with a safe return in the fall is vaccination of as many people as possible. Another is incorporating genomic surveillance of SARS-CoV-2 so we’re able to track the spread of the virus,” Azarian says. “What we want to do is build a ring of protection around the students and employees at the school so that basically we’re identifying cases as early as possible and implementing public health measures. We’ll also be able to assess the effectiveness of those measures using viral sequencing data.”
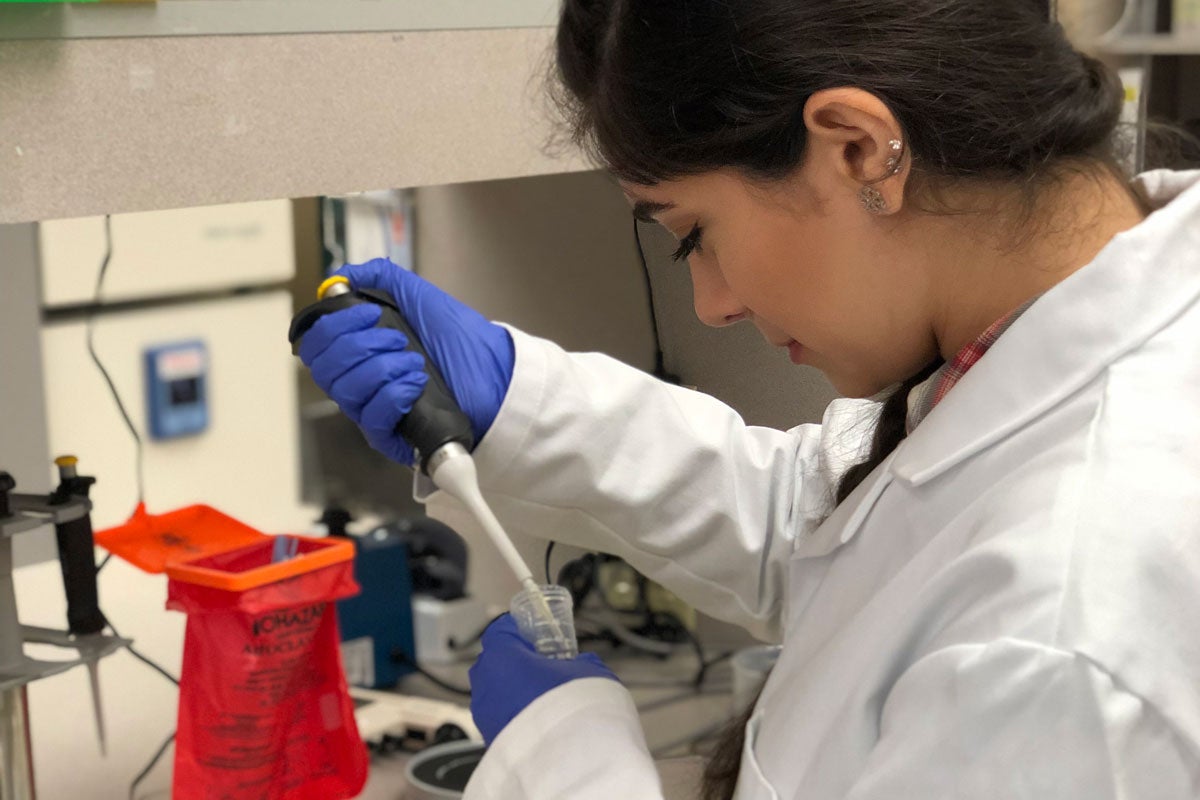
Azarian’s lab takes samples — with individuals’ identities redacted — from positive COVID-19 tests and isolates the virus’ RNA, or genetic code. The samples are obtained from random testing on campus as well as symptomatic patients at the Student Health Center and the Parking Garage A testing site.
The lab prepares the RNA sample and then inputs it into a genome sequencing platform that enables hundreds of samples to be simultaneously analyzed with the aid of the high-performance computing cluster at UCF. This allows the lab to compare viral sequences among individuals with COVID-19 to look for similarities or differences.
For instance, Azarian says, this analysis could determine if a set of cases is linked to one residence hall, or a social event, which would help officials identify additional cases and determine appropriate public health interventions.
“I like to describe this as building family trees of pathogens,” he says. “By looking at the relatedness between strains, we can infer a number of things — like how fast an epidemic is spreading, how fast it is evolving and whether it is developing resistance to any of our interventions such as vaccines.”
“By looking at the relatedness between strains, we can infer a number of things — like how fast an epidemic is spreading, how fast it is evolving and whether it is developing resistance to any of our interventions such as vaccines.” — Taj Azarian, assistant professor
Azarian says he was drawn to working at UCF because of its genomics and bioinformatics faculty cluster, which was created to inspire cross-cutting research that leverages UCF’s strengths in biomedical sciences, evolution and ecology, and computer science. That faculty cluster has played a role in aiding his genomic surveillance work during the pandemic.
Since the beginning of the pandemic, he has been working with the Florida Department of Health to monitor the emergence and spread to COVID-19 in the community.
Now his focus will shift to doing the same for UCF, where his lab will be able to compare viral sequences from UCF to those collected throughout Florida and abroad.
It’s part of a broader initiative led by the Centers for Disease Control and Prevention, which works closely with researchers and public health labs in the United States to generate, share and analyze viral sequencing data. Researchers and public health officials can analyze and compare the data in a larger context to better understand the virus, detect a potential outbreaks of related cases, develop interventions, and monitor emerging variants. Recently, Azarian and a colleague at the University of Florida were appointed to collaborate on a project funded by The Rockefeller Foundation to become part of a U.S. Regional Accelerators for Genomic Surveillance program.
Monitoring variants has become increasingly crucial due to their potential to spread easier or cause more severe disease. Azarian says that there is a growing concern for variants for which the vaccine is less effective.
Azarian emphasizes that ensuring individuals’ privacy and personal information through the process is a priority and that no one’s DNA is retained anywhere.
“We’re only interested in the RNA of the virus,” he says. “In any of the work that we do, we are not looking for the human DNA that may be present in the sample. We’re only focusing on the virus’ sequence, and once we perform the viral RNA extraction, those samples are discarded. Furthermore, when our results are uploaded to the online repository, we do a screening to make sure there is no human DNA data being uploaded.”
Azarian’s work in his lab is assisted by two undergraduate students, a recent graduate and two post-doctoral fellows at UCF.
“As an aspiring physician, being involved in such a project is an honor and a once-in-a-lifetime opportunity,” says Anita Samadabadi ’20, a Burnett Honors Scholar who graduated in December with a bachelor’s degree in biomedical sciences. “This opporutnity allows me to apply what I learned in the classroom about biomedical research to real-world scenarios. My hope is that we all will learn valuable lessons from the COVID-19 pandemic so we never have to face another pandemic again in the future. But if that unfortunately happens, I am sure that the lessons and experiences I am acquiring by being part of this project will help me be an asset for my community.”
Azarian says this field will continue to be critical even as more of the world’s population is vaccinated.
“We as a community are starting to see the light at the end of the tunnel because of the vaccine, and I think the No. 1 question on everyone’s mind is, ‘Did we win the fight?’” he says.
“Right now, I feel like we are in a race between getting people immunized and the spread of some of these new variants. Even as immunization rates are climbing, it’s imperative we continue this surveillance so we can identity whether those variants continue to emerge and what that means for the future of the vaccine — whether it will need to be updated at some point to be more like the seasonal influenza vaccine, or if this one is going to work against the strains now and the ones that may emerge in the future. Genomic surveillance will help us answer these questions.”